K-mer analysis and genome size estimate
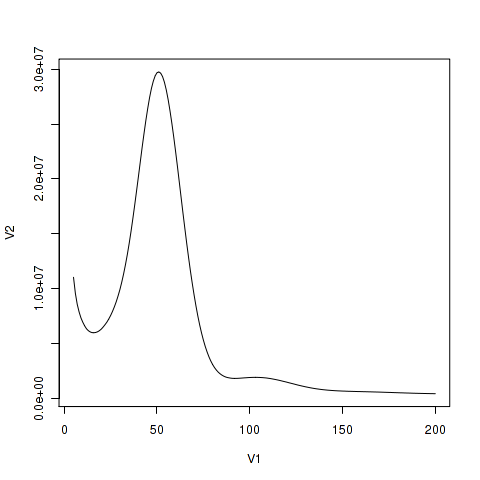

Plants, Free Full-Text
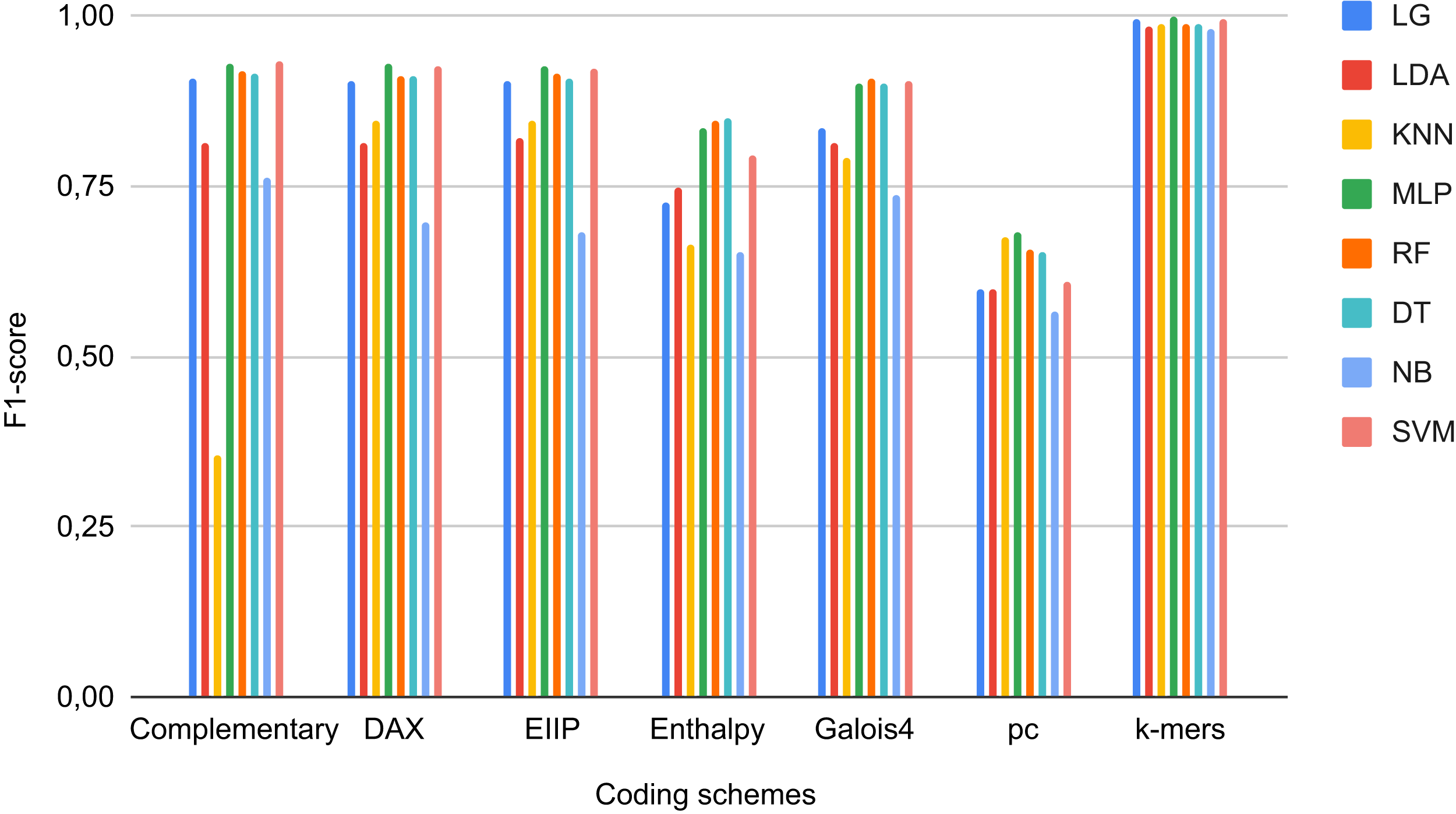
K-mer-based machine learning method to classify LTR-retrotransposons in plant genomes [PeerJ]

significant differences of genome size estimates when varying the Max kmer coverage · Issue #21 · schatzlab/genomescope · GitHub

Workflow (Illumina Inc), Bioz
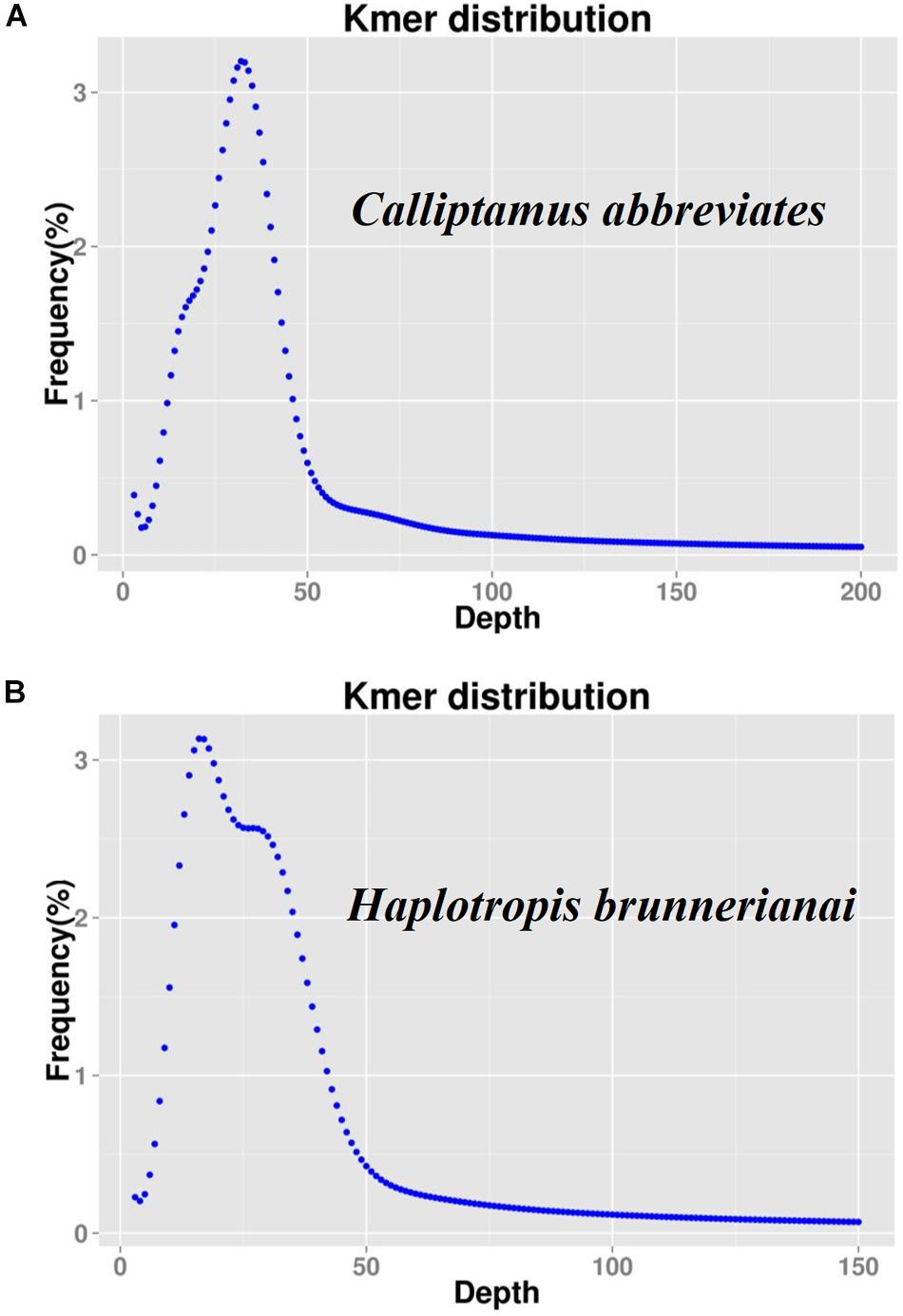
Frontiers Genome Size of 17 Species From Caelifera (Orthoptera) and Determination of Internal Standards With Very Large Genome Size in Insecta

De novo whole-genome assembly of a wild type
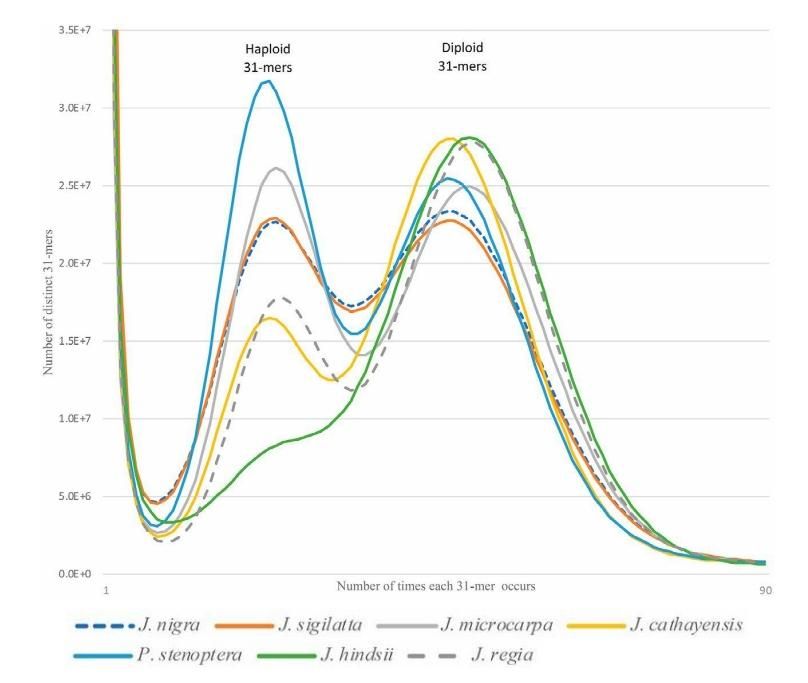
UCD Bioinformatics Core Workshop

k-mer - Wikipedia

Unbiased K-mer Analysis Reveals Changes in Copy Number of Highly Repetitive Sequences During Maize Domestication and Improvement

Merqury: reference-free quality, completeness, and phasing assessment for genome assemblies, Genome Biology
Large-scale k-mer-based analysis of the informational properties of genomes, comparative genomics and taxonomy

K-mer analysis of three Tokudaia species genomes. (a) T. osimensis (b)

K-Mer-Based Genome Size Estimation in Theory and Practice
Calculate the N50 for the example contigs below

K-mer (k = 17) analysis for estimating the genome size of R.